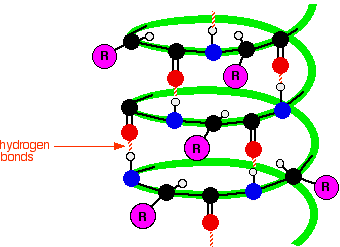
Sequence determinants of alpha helical folding and unfolding You really need space-filling models to be able to visualize that. One limitation of the paper is that with just 2D images and words, it is very difficult to convey exactly what drives the differences in multimerization. Sadly, 22 years later, this work remains “cutting edge” in the sense that we still don’t understand any better why each residue has such different effects on multimerization. These dramatic differences showed that specific residues - and not just the hydrophobicity of the interaction surface - mattered a lot. When V was involved, you got a mixture of species. but when a was all L and d was I, you got a tetramer. 
When a was all I and d was all L, you got a dimer. The authors created a series of mutants of the GCN4 coiled coil, changing the residues in the a and d positions either to all V, all I, all L, or various pairwise combinations. Helices are almost never seen naked and do not have charged residues on both sides of the helix - instead, they have one hydrophobic face. Their designed peptide was very different from any peptide seen in living organisms. Helix formation was optimal with E and K in the i±4n ridge, and this suggested that the formation of salt bridges, and the dipole moment of the protein, were important. They also discuss a de novo designed peptide consisting of all alanines except for repeating units of E and K organized along i±4n or i☓n ridges. They review the experiments of Baldwin’s group with RNAse A C-peptide, which are covered in more detail in the lecture notes below. This chapter was written in 1990, when hopes were high that de novo design of peptides was just around the corner. This book chapter chronicles some of the landmark experiments and observations which informed our understanding of how sequence drives alpha helix formation. They found that it was necessary that near the ends there be side chains which can satisfy some of the hydrogen bonding requirements of the backbone, and then another residue which can shield those side chains. They took alpha helical structures and modeled different amino acid substitutions at the N and C termini in silico. This is consistent with the dipole moment model of alpha helix formation.Ī related and interesting set of analyses were done by. D and E, the negatively charged / acidic residues, are preferred near the N terminus K and R, the positively charged / basic residues are preferred near the C terminus. Proline is the amino acid most rarely seen in alpha helices, for two reasons: 1) it cannot rotate around its N-C bond, and 2) its N is not protonated, so it cannot participate in the hydrogen bonding that defines the alpha helix backbone. This model was preferred because it gave stronger signals of preference for which amino acid should be the cap. Instead, they chose a model where the last amino acid whose alpha carbon (αC) falls within the cylinder defined by the helix, is considered to be the N-cap or C-cap, meaning the first or last residue in the helix. A model based on φ and ψ angles was rejected. They considered two possible models for defining where a helix starts and stops. Richardson & Richardson 1988Īt the time, figuring out where you define the start and stop of an alpha helix was a major problem. The readings for this week are chapter 15 of Branden & Tooze on immunoglobulins, and. These are my notes from week 4 of MIT course 7.88j: Protein Folding and Human Disease, held by Dr. Protein folding 04: Formation of alpha helices
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
